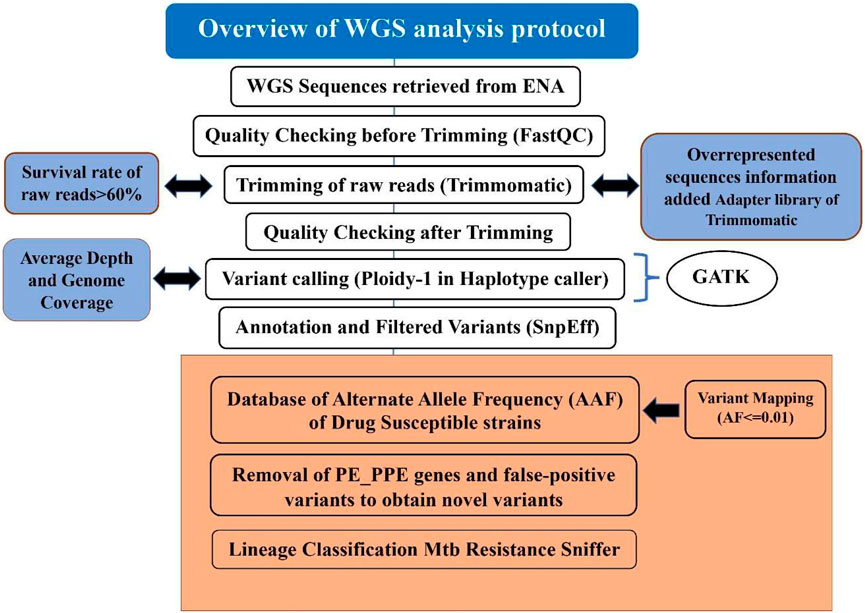
Frontiers | MycoVarP: Mycobacterium Variant and Drug Resistance Prediction Pipeline for Whole-Genome Sequence Data Analysis

A Bioinformatics Whole-Genome Sequencing Workflow for Clinical Mycobacterium tuberculosis Complex Isolate Analysis, Validated Using a Reference Collection Extensively Characterized with Conventional Methods and In Silico Approaches | Journal of ...
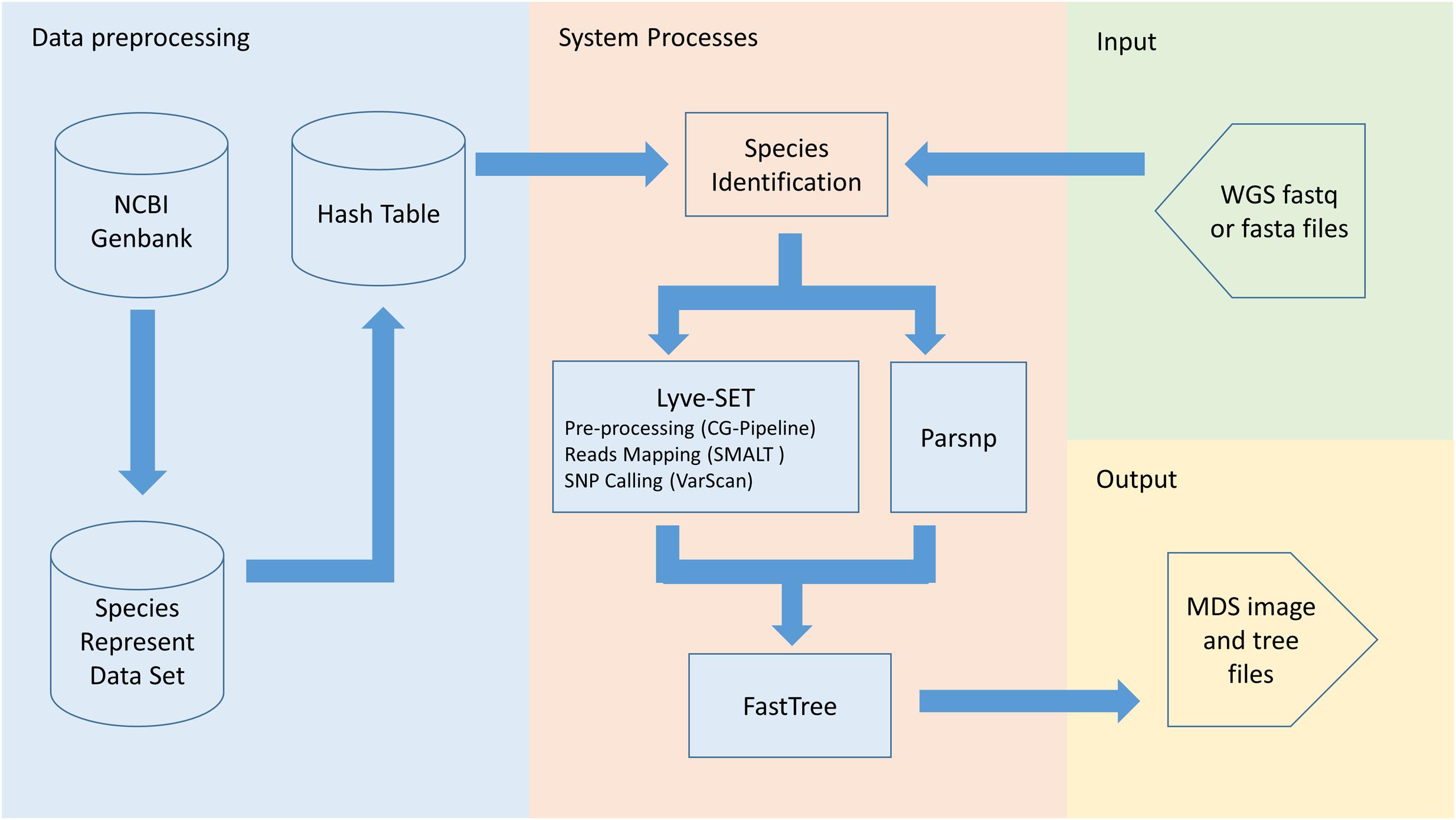
Frontiers | PathoBacTyper: A Web Server for Pathogenic Bacteria Identification and Molecular Genotyping
![sv-callers: a highly portable parallel workflow for structural variant detection in whole-genome sequence data [PeerJ] sv-callers: a highly portable parallel workflow for structural variant detection in whole-genome sequence data [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/8214/1/fig-1-2x.jpg)
sv-callers: a highly portable parallel workflow for structural variant detection in whole-genome sequence data [PeerJ]

Whole-Genome Sequencing of Bacterial Pathogens: the Future of Nosocomial Outbreak Analysis | Clinical Microbiology Reviews
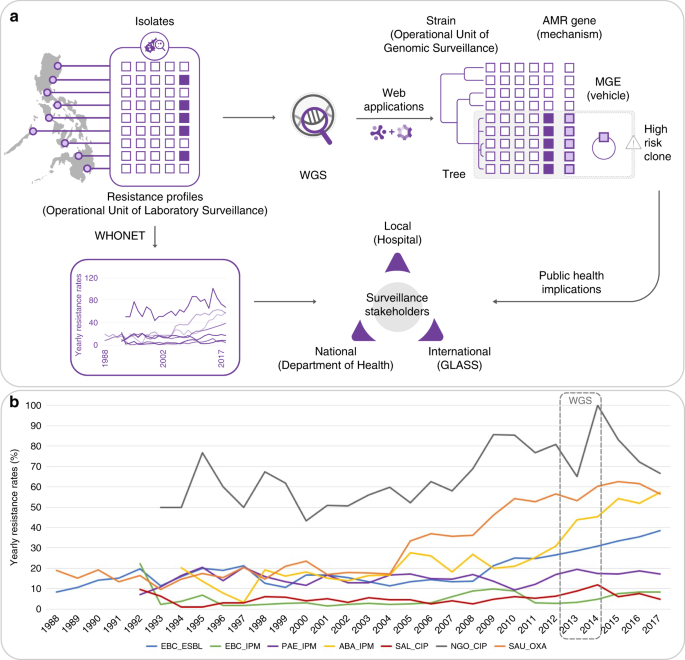
Integrating whole-genome sequencing within the National Antimicrobial Resistance Surveillance Program in the Philippines | Nature Communications

PDF) Leveraging TOPMed Imputation Server and Constructing a Cohort-Specific Imputation Reference Panel to Enhance Genotype Imputation among Cystic Fibrosis Patients

Whole genome sequencing delineates regulatory, copy number, and cryptic splice variants in early onset cardiomyopathy | npj Genomic Medicine

Workshop KEK - CC-IN2P3 KEK new Grid system 27 – 29 Oct. CC-IN2P3, Lyon, France Day2 14: :55 (40min) Koichi Murakami, KEK/CRC. - ppt download

A Bioinformatics Whole-Genome Sequencing Workflow for Clinical Mycobacterium tuberculosis Complex Isolate Analysis, Validated Using a Reference Collection Extensively Characterized with Conventional Methods and In Silico Approaches | Journal of ...
![PDF] Evaluation of WGS-subtyping methods for epidemiological surveillance of foodborne salmonellosis PDF] Evaluation of WGS-subtyping methods for epidemiological surveillance of foodborne salmonellosis](https://i1.rgstatic.net/publication/342717659_Evaluation_of_WGS-subtyping_methods_for_epidemiological_surveillance_of_foodborne_salmonellosis/links/5f0358eaa6fdcc4ca44ebc81/largepreview.png)
PDF] Evaluation of WGS-subtyping methods for epidemiological surveillance of foodborne salmonellosis
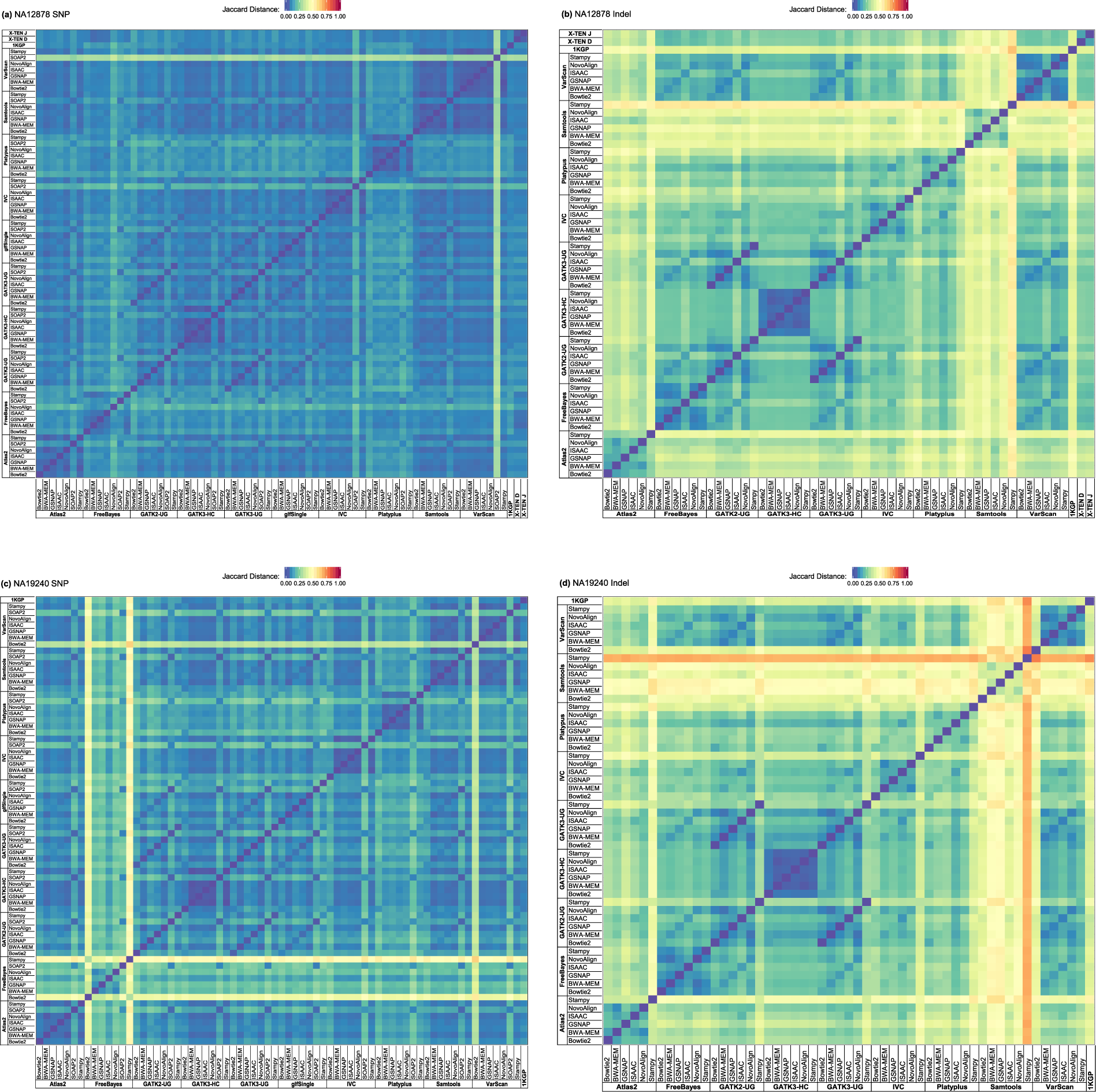
Comparative analysis of whole-genome sequencing pipelines to minimize false negative findings | Scientific Reports
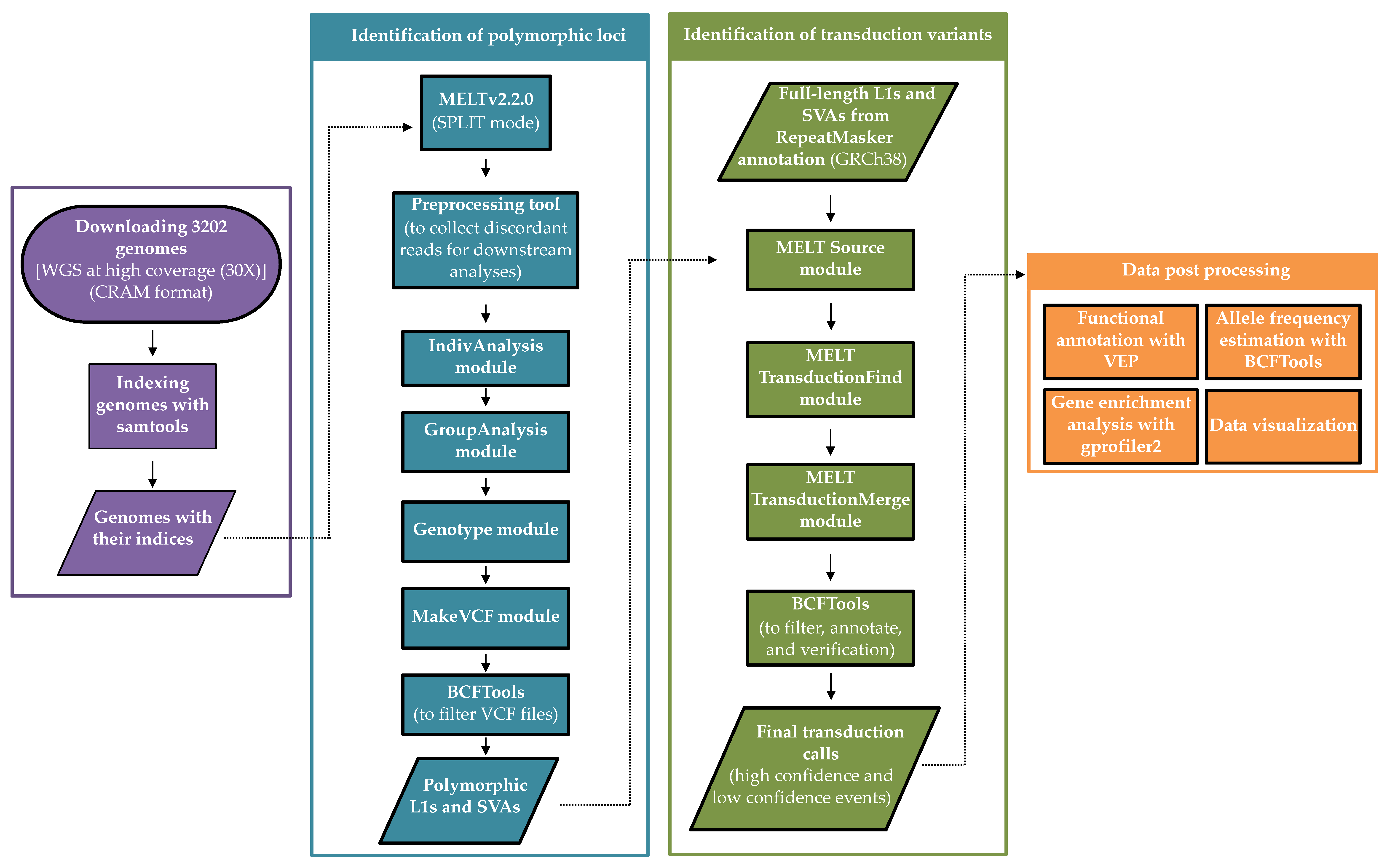
Biology | Free Full-Text | A Map of 3′ DNA Transduction Variants Mediated by Non-LTR Retroelements on 3202 Human Genomes | HTML
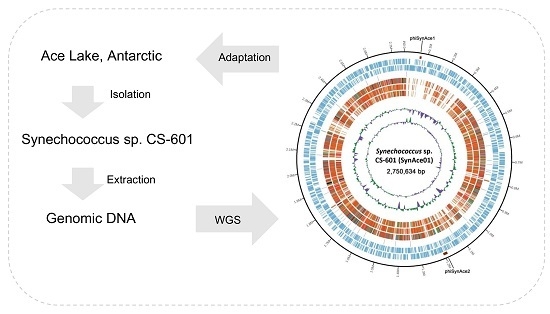
IJMS | Free Full-Text | Complete Genome Sequence and Comparative Analysis of Synechococcus sp. CS-601 (SynAce01), a Cold-Adapted Cyanobacterium from an Oligotrophic Antarctic Habitat | HTML